Multiple Sequence Alignment (MSA) with Genetic Algorithm + Qwen2.5
GA-based MSA with PyTorch GPU fitness and local LLM JSON analysis
- Implemented a lightweight genetic-algorithm–based multiple sequence alignment (MSA) on Kaggle’s Sequence Alignment (Bioinformatics) Dataset.
- Integrated affine gap penalties (PAM250/BLOSUM62), tournament selection, residue-count–preserving crossover, and improving mutations.
- Accelerated fitness evaluation with PyTorch on GPU (sum-of-pairs with affine gaps), plus elitism and immigration per generation.
- Exported best alignments in CLUSTAL (.aln) and FASTA (.fasta) formats.
- Computed per-column conservation (entropy), gap density, and pairwise % identity; visualized with heatmaps and dendrograms.
- Enhanced interpretability with a local LLM (Qwen2.5-1.5B via llama-cpp, no API keys), producing validated JSON reports of conserved blocks, gap clusters, and closest/divergent sequence pairs.
Repo: Github
Dataset: Kaggle Dataset
Model: Qwen2.5-1.5B-Instruct (GGUF)
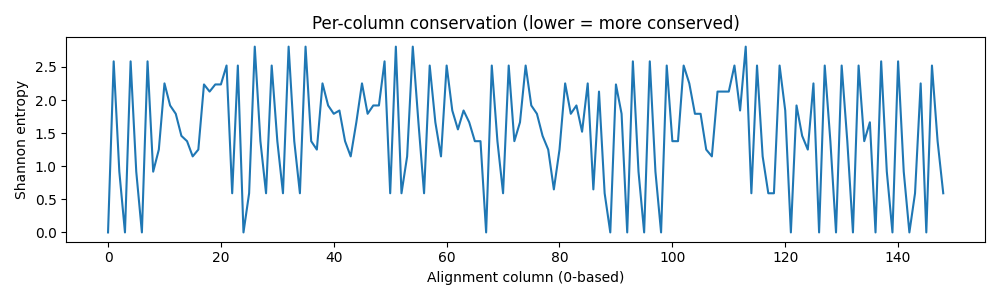
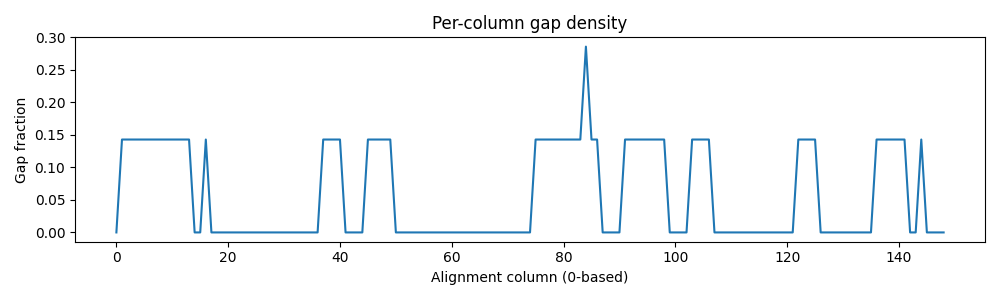
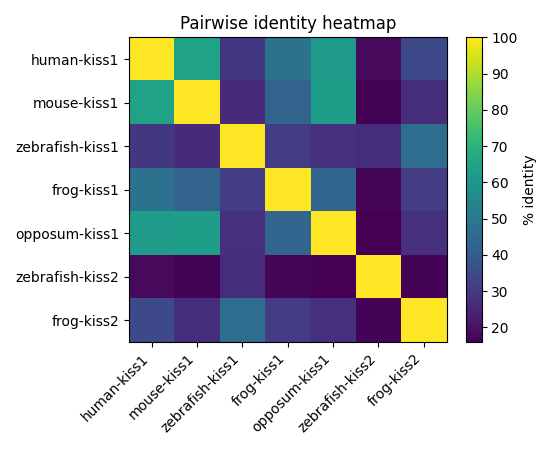
Left (top): Column-wise conservation (entropy). Left (bottom): Gap density across alignment. Right: Pairwise % identity heatmap of aligned sequences.
Highlights
- Frameworks: PyTorch, Biopython, matplotlib, llama-cpp-python
- GA features: Affine gaps, tournament selection, residue-preserving crossover, improving mutations, elitism, immigration
- Outputs: CLUSTAL/FASTA alignments, entropy & gap plots, identity heatmap, UPGMA dendrogram, JSON report
- LLM integration: Qwen2.5-1.5B (local, no API) for automated alignment summaries